Actually, the first thing I would say is this: use tools you are familiar with. If it’s Python, great, use it. And also, use the best tool for the job. If it’s Python, great, use it. And also, it’s okay to use a tool for one task because you’re already using it for all sorts of other tasks and so you have it. If you’re pounding nails all day it’s fine if you’re also using the hammer to open a beer bottle or scratch your back. Similarly, if you are programming in Python all day long then it is fine if you are also using it to fit mixed linear models. If it works for you, great! keep going. But if you’re struggling, if things seem harder than expected, this article series may be for you.
I think people over-index Python. Language for data science. It has some limitations that I think are quite notable. There are many data-science tasks that I would prefer to do in R than in Python. I believe the reason Python is so widely used in data science is a historical accident, as well, rather than an expression of its inherent suitability for data-science work right now at most things.
Also, I think Python is quite good for deep learning. This is why PyTorch is the industry standard. When I’m talking about data science here, I’m specifically excluding deep learning. I’m talking about all the other things: data perturbation, exploratory data analysis, visualization, statistical modeling, etc. And, as I said in my opening paragraph, I understand that if you’re already working in Python all day for a good reason (e.g., training an AI model) then you might want to do everything else in Python as well. I’m doing this myself, in the deep learning classes I teach. That doesn’t mean I can’t be frustrated by how cumbersome data science often is in the Python world.
Let’s start with my lived experience, without giving any explanation as to what might be causing it. I have been running a research lab in computational biology for more than two decades. During this time I have worked with approximately thirty graduate students and postdocs, all very capable and accomplished computational scientists. The policy in my lab is that everyone is free to use any programming language and tools. I don’t tell people what to do. And often, people choose Python as their programming language of choice.
So here’s a typical experience I typically have with students who use Python. A student comes to my office and shows me some results. I say “That’s great, but can you quickly plot the data in this other way?” or “Can you quickly calculate this quantity I created and tell me what it looks like when you plot it?” or similar. Usually, the requests I make are for something I know I can do in R in a few minutes. Examples include converting a boxplot to a violin or vice versa, converting a line plot to a heatmap, plotting density estimates instead of histograms, performing calculations on ranked data values instead of raw data values, etc. Without fail, the response from students who use Python is: “It’s going to take me a while. Let me sit at my desk and figure it out and then I’ll come back.” Now let me be very clear: these are strong students. The issue isn’t that my students don’t know their tools. It seems to me that this is a problem with the devices themselves. They seem so burdensome or confusing that requests that I think should be trivial often aren’t.
Whatever the reason for this experience, I have to conclude that there is something fundamentally wrong with how data analysis works in Python. This may be a problem of the language itself, or simply the limits of the software libraries available, or a combination thereof, but whatever the case, the effects are real and I see them regularly. In fact, I have another example, if you’re tempted to counter, “It’s a skills issue; get better students.” Last time, I taught a class on AI models for biology with an experienced data scientist who does all his work in Python. He knows NumPy and pandas and matplotlib like the back of his hand. In class, I covered all the theory, and he covered classroom exercises in Python. So I got a chance to watch an expert in Python work through several examples. And my reaction to the code examples was frequently, “Why is this so complicated?” Many times, what I thought would be a few lines of simple R code turned out to be quite long and quite complex. I certainly couldn’t have written that code without extensive study and without fully preparing my brain in terms of using programming patterns. It felt very strange, but not in the sense of “wow, this is so different but also so beautiful” but in the sense of “wow, this is so different and weird and cumbersome.” And again, I don’t think it’s because my coworker isn’t very good at what he’s doing. He is very good. The problem appears to lie in the basic structure of the devices.
Let’s step back for a moment and discuss some basic considerations for choosing a language for data science. When I say data science, I mean dissecting and summarizing data, finding patterns, fitting models, and creating visualizations. In short, this is the kind of stuff scientists and other researchers Do this while they are analyzing their data. This activity is separate from data engineering or application development, even if the application performs data-heavy workloads.
Data science, as I define it here, involves lots of interactive exploration of data and quick one-off analyzes or experiments. Therefore, any language suitable for data science must be interpreted in an interactive shell or notebook format. This also means that performance considerations are secondary. When you want to do quick linear regression on some data you’re working with, you don’t care whether the task will take 50 milliseconds or 500 milliseconds. What you care about is whether you can open a shell, type a few lines of code, and get results in a minute or two, versus setting up a new project, writing all the boilerplate to please the compiler, and then spending more time compiling your code than running it.
If we accept that being able to work interactively and with low startup-costs is an important feature of a language for data science, we immediately turn to scripting languages such as Python, or data-science specific languages such as R or Matlab or Mathematica. There’s also Julia, but honestly I don’t know enough about it to write coherently about it. What I do know is that it is probably the best data science language out there. But I have noted that some people Those who have used it extensively have their doubts. Anyhow, I will not discuss it further here. I also wouldn’t consider proprietary languages like Matlab or Mathematica, or fairly obscure languages that lack a broad ecosystem of useful packages like Octave. This leaves us with R and Python as realistic options to consider.
Before continuing, I’d like to offer a few more thoughts about the display. Performance generally coincides with other features of a language. In simple terms, performance comes at the expense of either additional overhead for the programmer (as in Rust) or increased risk of obscure bugs (as in C) or both. For data science applications, I do not consider obscure bugs or high risk of incorrect results to be acceptable, and I also think that convenience for programmers is more important than raw performance. Computers are fast and it hurts to think. I would prefer to spend less mental energy telling the computer what to do and wait a little longer for results. Therefore, the easier a language can make my work, the better it will be. If I’m really performance-limited in an analysis, I can always rewrite that particular part of the analysis in Rust, once I know what I’m doing and what calculations I need.
An important component of not making my job more difficult than necessary is separating the logic of the analysis from the logistics. What I mean by this is that I want to be able to specify at a conceptual level how the data should be analyzed and what the result of the calculations should be, and I don’t want to think about the logistics of how the calculations are done. As a general rule, if I have to think about data types, numerical indexes, or loops, or if I have to manually disassemble and reassemble a dataset, chances are I’ll get stuck in the logistics.
To provide a concrete example, consider the dataset Penguins from the Palmer Archipelago. There are three different penguin species in the dataset, and the penguins live on three different islands. Suppose I want to calculate the mean and standard deviation of penguin weight for each combination of penguin species and island, excluding any cases where the penguin’s body weight is not known. An ideal data science language would allow me to express this calculation in these words, and it would require about the same amount of code as it took me to write this sentence in the English language. And actually this is possible in both R and Python.
Here’s the relevant code in R, using the Tidyvers approach:
library(tidyverse)
library(palmerpenguins)
penguins |>
filter(!is.na(body_mass_g)) |>
group_by(species, island) |>
summarize(
body_weight_mean = mean(body_mass_g),
body_weight_sd = sd(body_mass_g)
)And here is the equivalent code in Python using the pandas package:
import pandas as pd
from palmerpenguins import load_penguins
penguins = load_penguins()
(penguins
.dropna(subset=['body_mass_g'])
.groupby(['species', 'island'])
.agg(
body_weight_mean=('body_mass_g', 'mean'),
body_weight_sd=('body_mass_g', 'std')
)
.reset_index()
)These two examples are very similar. At this level of analysis complexity, Python works fine. I would consider the R code a little easier to read (note how many quotes and brackets the Python code requires), but the differences are minor. In both cases, we take the penguin dataset, remove those penguins whose body weight is missing, then specify that we want to calculate separately on each combination of penguin species and island, and then calculate the means and standard deviations.
Compare this to the equivalent code that’s full of logistics, where I’m using only basic Python language features and no special data wrangling packages:
from palmerpenguins import load_penguins
import math
penguins = load_penguins()
# Convert DataFrame to list of dictionaries
penguins_list = penguins.to_dict('records')
# Filter out rows where body_mass_g is missing
filtered = [row for row in penguins_list if not math.isnan(row['body_mass_g'])]
# Group by species and island
groups = {}
for row in filtered:
key = (row['species'], row['island'])
if key not in groups:
groups[key] = []
groups[key].append(row['body_mass_g'])
# Calculate mean and standard deviation for each group
results = []
for (species, island), values in groups.items():
n = len(values)
# Calculate mean
mean = sum(values) / n
# Calculate standard deviation
variance = sum((x - mean) ** 2 for x in values) / (n - 1)
std_dev = math.sqrt(variance)
results.append({
'species': species,
'island': island,
'body_weight_mean': mean,
'body_weight_sd': std_dev
})
# Sort results to match order used by pandas
results.sort(key=lambda x: (x['species'], x['island']))
# Print results
for result in results:
print(f"{result['species']:10} {result['island']:10} "
f"Mean: {result['body_weight_mean']:7.2f} g, "
f"SD: {result['body_weight_sd']:6.2f} g")This code is very long, has many loops, and it obviously pulls the dataset apart and then puts it back together again. Regardless of language choice, I hope you can see that the version without logistics is better than the version that gets bogged down in logistics details.
I’ll end things here for now. This post is quite long. In future installments, I will discuss specific issues that make data analysis more complex in Python than in R. In short, I believe there are many reasons why Python code often turns to dealing with data logistics. As much as a programmer may try to avoid the logistics and stick to high-level conceptual programming patterns, either the language or the available libraries get in the way and thwart those efforts. I will go into details soon. Stay tuned.
LLMs are great at programming—how can they be so bad at it?

Despite the overall hype in all things AI, especially among the tech crowd, we have yet to see much in terms of product-market fit and actual commercial success for AI – or more specifically, LLM – outside a fairly narrow range of application areas. Apart from chatty chatbots, AI girlfriends and maybe efficient document searching, the main applications…
No, AlphaFold has not completely solved protein folding
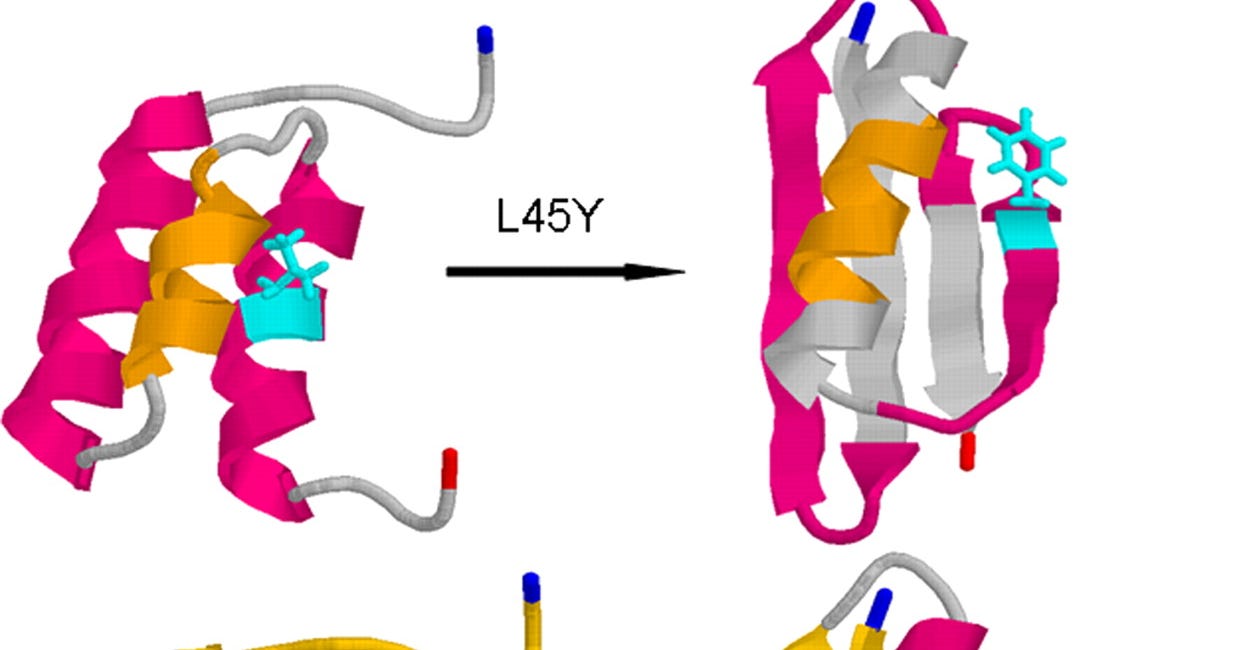
AlphaFold has captured the imagination of people outside biology to a degree not usually seen for a technical tool of computational biology. No tech bro in Silicon Valley has any opinion on HMMER, BLAST, or FoldX, or their potential impact on the future of humanity. But when it comes to
<a href