Figure 2.1: Statistical Process Control!
In this workshop, we will learn how to do statistical process control in Python using statistical tools plotnine Visualization! Statistical process control refers to (1) using statistics to measure variations in product quality over time and (2) identifying benchmarks to know when intervention is needed. let’s get started!
launch
packages
# Remember to install these packages using a terminal, if you haven't already!
!pip install pandas plotnine scipywe will experiment pandas For data manipulation, plotnine For visualization, and scipy For statistical purposes.
import pandas as pd
from plotnine import *custom work
This workshop uses custom functions functions/ Directory. You may need both:- functions_distributions.py – For reliability and delivery functions – functions_process_control.py – for statistical process control tasks
To use these functions, you need to get them from the repository at github.com/timothyfraser/sigma/tree/main/functions.
Add functions directory to your python path
import sys
import os
# Add the functions directory to Python path
sys.path.append('functions') # or path to wherever you placed the functions folderOnce you have the functions available, you can import them:
from functions_distributions import density, tidy_density, approxfun
# from functions_process_control import ggprocess, ggsubgroup, ggmoving, ggcapability # if neededour case
For today’s workshop, we will consider why quality control matters in a local economy by examining the case of the Japanese hot springs bath economy! hot springs, or onsenHot springs are a major source of tourism and recreation for families in Japan, drawing residents from across the country each year, often to rural communities where the perfect geological conditions produce naturally occurring hot springs. Restaurants, taxi and bus companies, and many service sector companies rely on their local efforts to bring a steady stream of tourists into the local economy. Therefore, it is often in the best interest onsen To ensure quality control, operators must monitor the temperature, minerals, or other aspects of their hot spring baths in order to maintain their company’s (and city’s!) reputation for quality rest and relaxation!
Onsen-Those who visit often seek Specific Types of Hot Springs, Why It’s Important onsen Really delivering what it advertises! Sarbulia and Payyappallimana (2012) describe some of these benchmarks.
- temperature: Onsen are divided into “additional hot springs” (
>42°C), “hot Springs” (41~34°C), and “Warm Springs” (33~25°C, - pH: Onsen is classified as “acidic” (
pH < 3), “mildly acidic” (pH 3~6), “neutral” (pH 6~7.5), “mild alkaline” (pH 7.5~8.5), and “alkaline” (pH > 8.5, -
sulfur:sulphur onsen There is usually about 2 mg of sulfur per 1 kg of hot spring water; sulfur levels Sure More than 1 mg counts as sulfur onsen. (It smells like rotten eggs!)
These are good examples of quality control metrics onsen Operators might want to keep an eye out!
](https://timothyfraser.com/sigma/images/9_monkeys.png)
Figure 4.1: Monkeys are onsen fans too! Reading more here!
our data
You have been assigned to evaluate quality control at the local level onsen In sunny Kagoshima Prefecture! Every month, for 15 months, you systematically took 20 random samples of hot spring water and recorded it temperature, pHAnd sulfur level. How can you determine that this onsen There is a danger of moving out of one sector of the market (e.g. extra hot!) into another (just normal hot springs?).
Let’s read into our data workshops/onsen.csv,
# Add functions directory to path if not already there
import sys
if 'functions' not in sys.path:
sys.path.append('functions')
from functions_distributions import density, tidy_density, approxfun
water = pd.read_csv('workshops/onsen.csv')
water.head(3)## id time temp ph sulfur
## 0 1 1 43.2 5.1 0.0
## 1 2 1 45.3 4.8 0.4
## 2 3 1 45.5 6.2 0.9Process Descriptive Statistics
First, let’s understand our process by calculating some basic descriptive statistics. We will create a simple function to calculate the mean and standard deviation, which are fundamental to evaluating process variation.
from pandas import Series
def describe(x: Series):
x = Series(x)
out = pd.DataFrame({
'mean': [x.mean()],
'sd': [x.std()],
})
out['caption'] = ("Process Mean: " + out['mean'].round(2).astype(str) +
" | SD: " + out['sd'].round(2).astype(str))
return out
tab = describe(water['temp'])
tab## mean sd caption
## 0 44.85 1.989501 Process Mean: 44.85 | SD: 1.99Let’s now apply this to our temperature data to see the overall process mean and variation.
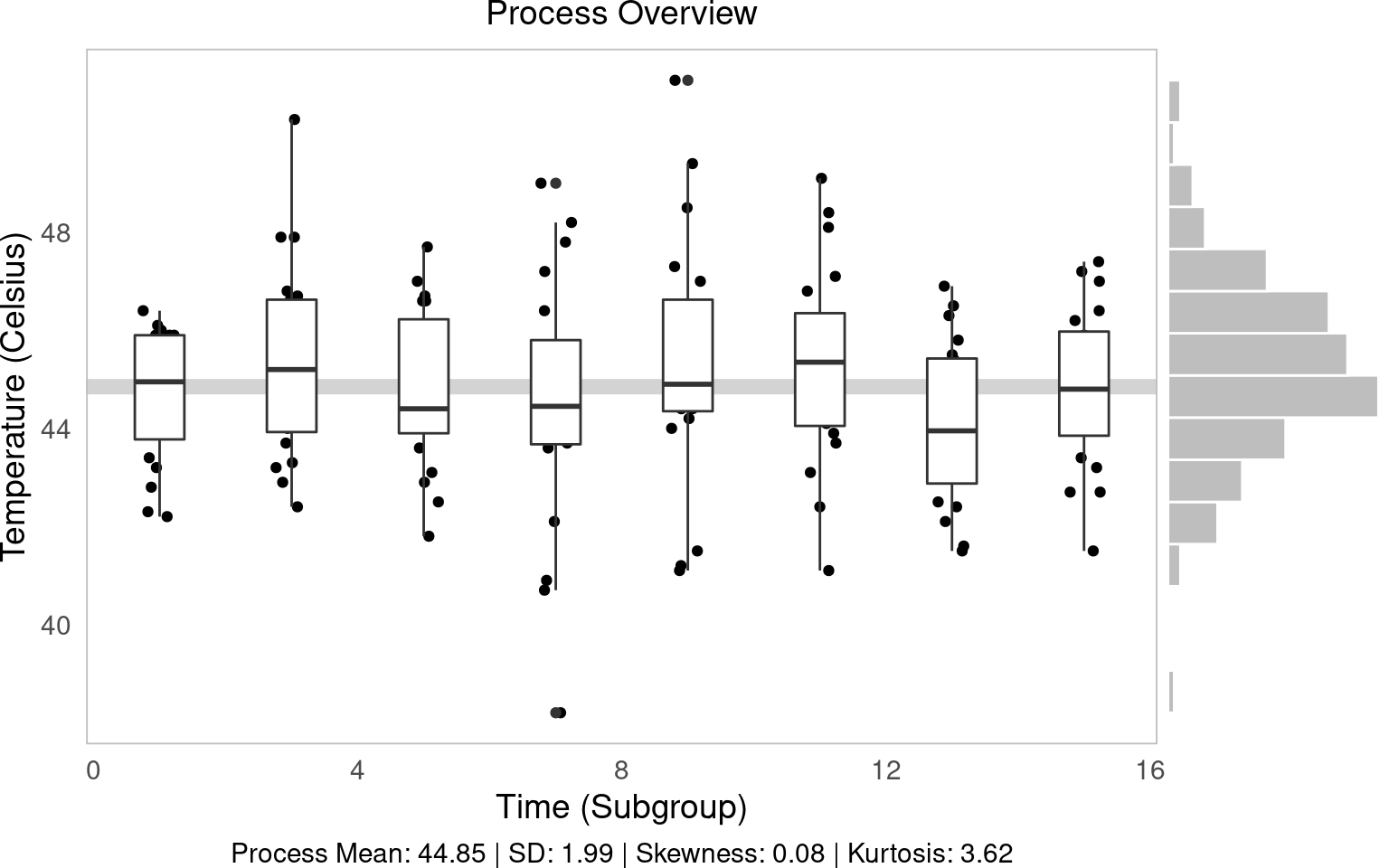
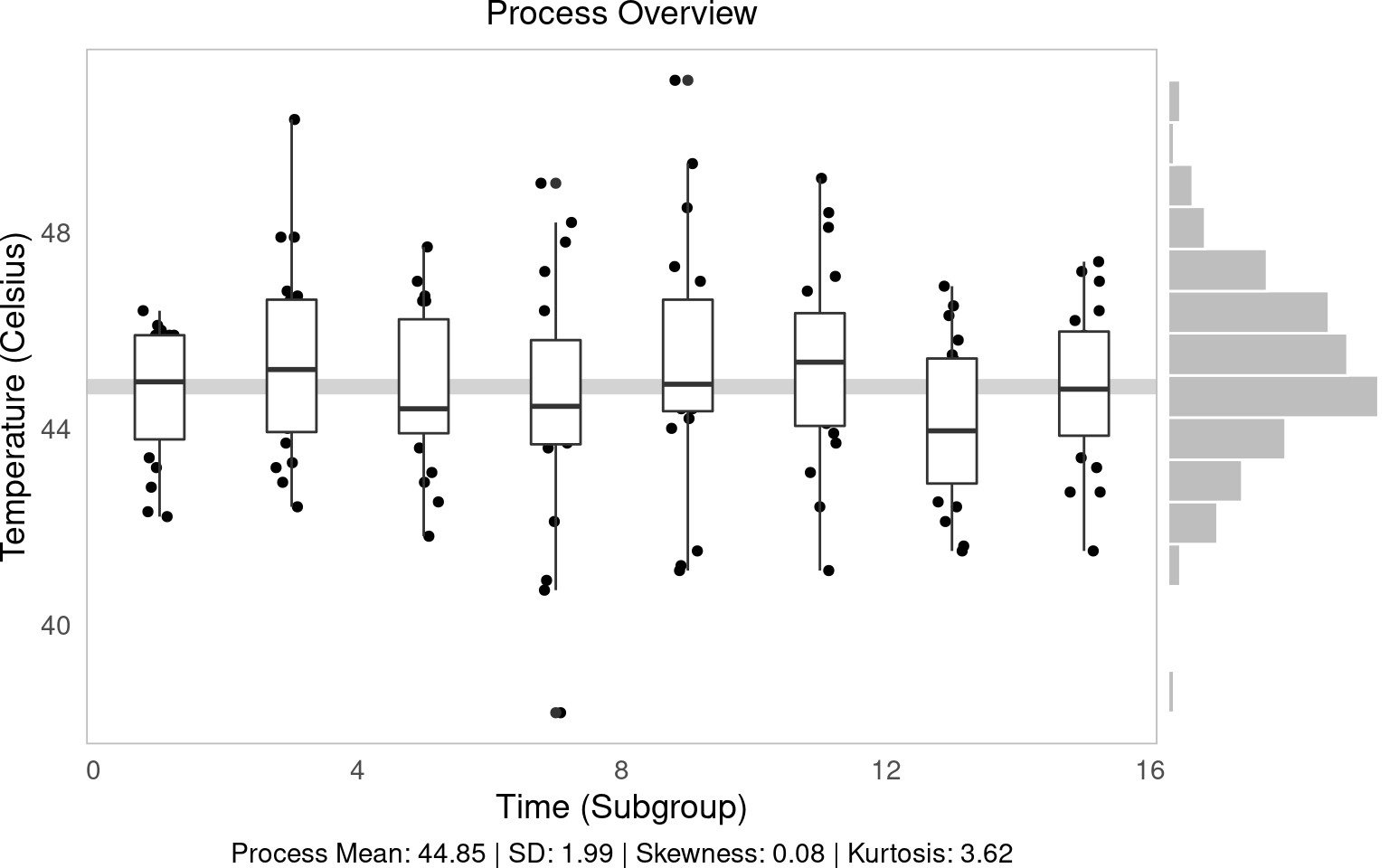
process overview view
The process overview chart is one of the most important tools in SPC. It shows us how our process behaves over time, helping us identify patterns, trends and potential issues. We will create a visualization that will show individual measurements, subgroup means, and overall process averages.
g1 = (ggplot(water, aes(x='time', y='temp', group='time')) +
geom_hline(aes(yintercept=water['temp'].mean()), color='lightgrey', size=3) +
geom_jitter(height=0, width=0.25) +
geom_boxplot() +
labs(x='Time (Subgroup)', y='Temperature (Celsius)', subtitle='Process Overview', caption=tab['caption'][0]))
# Save the plot
g1.save('images/05_process_overview.png', width=8, height=6, dpi=100)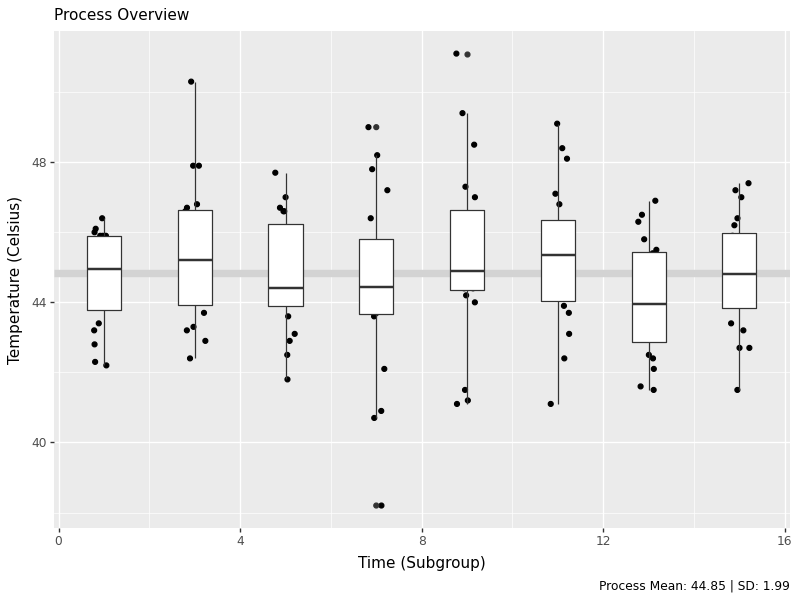
g2 = (ggplot(water, aes(x='temp')) + geom_histogram(bins=15, color='white', fill='grey') + theme_void() + coord_flip())
# Save the plot
g2.save('images/05_process_histogram.png', width=8, height=6, dpi=100)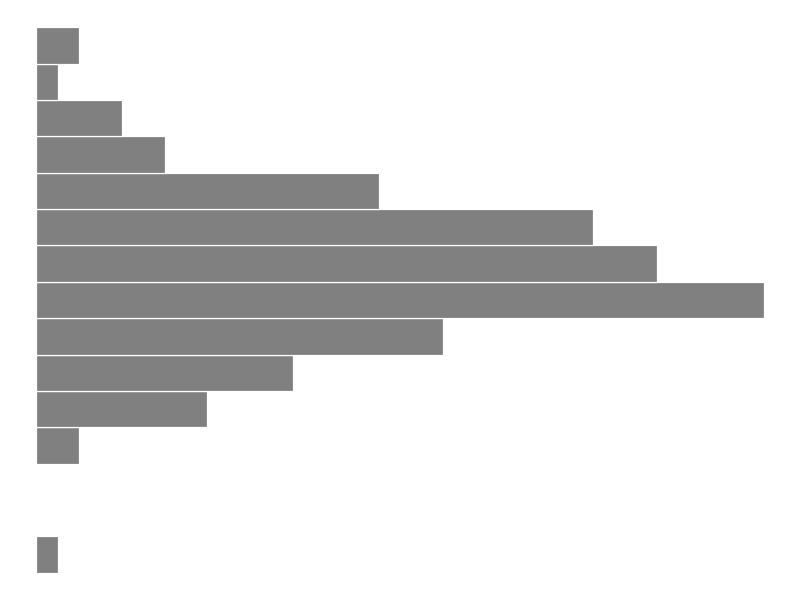
The histogram shows us the distribution of all temperature measurements, giving us information about overall process variation. This helps us understand if our process is focused and how much variation we are seeing.
Subgroup (within group) statistics
At SPC, we often work with subgroups – Small samples taken at regular intervals. This allows us to distinguish between common cause variation (inherent in the process) and special cause variation (due to specific events). Let’s calculate the statistics for each subgroup to see how the process behaves over time.
stat_s = (water.groupby('time').apply(lambda d: pd.Series({
'xbar': d['temp'].mean(),
'r': d['temp'].max() - d['temp'].min(),
'sd': d['temp'].std(),
'nw': len(d)
})).reset_index())
stat_s['df'] = stat_s['nw'] - 1
stat_s['sigma_s'] = ( (stat_s['df'] * (stat_s['sd']**2)).sum() / stat_s['df'].sum() )**0.5
stat_s['se'] = stat_s['sigma_s'] / (stat_s['nw']**0.5)
stat_s['upper'] = stat_s['xbar'].mean() + 3*stat_s['se']
stat_s['lower'] = stat_s['xbar'].mean() - 3*stat_s['se']
stat_s.head(3)## time xbar r sd nw df sigma_s se upper lower
## 0 1 44.635 4.2 1.342533 20.0 19.0 1.986174 0.444122 46.182366 43.517634
## 1 3 45.305 7.9 2.001440 20.0 19.0 1.986174 0.444122 46.182366 43.517634
## 2 5 44.765 5.9 1.628133 20.0 19.0 1.986174 0.444122 46.182366 43.517634Here we calculated the key statistics for each subgroup:
- xbar:mean of each subgroup
- R: Range (maximum – minimum) within each subgroup.
- sd: standard deviation within each subgroup
- sigma_s: clustered standard deviation within the subgroup.
- From: Standard error for each subgroup mean
Overall statistics (between groups)
Let’s now calculate overall process statistics that summarize behavior across all subgroups:
stat_t = pd.DataFrame({
'xbbar': [stat_s['xbar'].mean()],
'rbar': [stat_s['r'].mean()],
'sdbar': [stat_s['sd'].mean()],
'sigma_s': [(stat_s['sd']**2).mean()**0.5],
'sigma_t': [water['temp'].std()]
})
stat_t## xbbar rbar sdbar sigma_s sigma_t
## 0 44.85 7.2625 1.93619 1.986174 1.989501These data give us:
- xbbar: Grand mean (average of all subgroup means)
- rbar:average range in subgroups
- sdbar: average standard deviation across subgroups
- sigma_s: clustered standard deviation within the subgroup.
- sigma_t: total process standard deviation
Average and Standard Deviation Chart
Control charts are the heart of SPC. They help us monitor process stability over time and detect when a process goes out of control. We will create charts for both the subgroup mean (X-bar chart) and standard deviation (S chart).
labels = pd.DataFrame({
'time': [stat_s['time'].max()]*3,
'type': ['xbbar','upper','lower'],
'name': ['mean','+3 s','-3 s'],
'value': [stat_s['xbar'].mean(), stat_s['upper'].iloc[0], stat_s['lower'].iloc[0]]
})
control_chart = (ggplot(stat_s, aes(x='time', y='xbar')) +
geom_hline(aes(yintercept=stat_s['xbar'].mean()), color='lightgrey', size=3) +
geom_ribbon(aes(ymin='lower', ymax='upper'), fill='steelblue', alpha=0.2) +
geom_line(size=1) + geom_point(size=5) +
geom_label(data=labels, mapping=aes(x='time', y='value', label='name'), ha='right') +
labs(x='Time (Subgroups)', y='Average', subtitle='Average and Standard Deviation Chart'))
# Save the plot
control_chart.save('images/05_control_chart.png', width=8, height=6, dpi=100)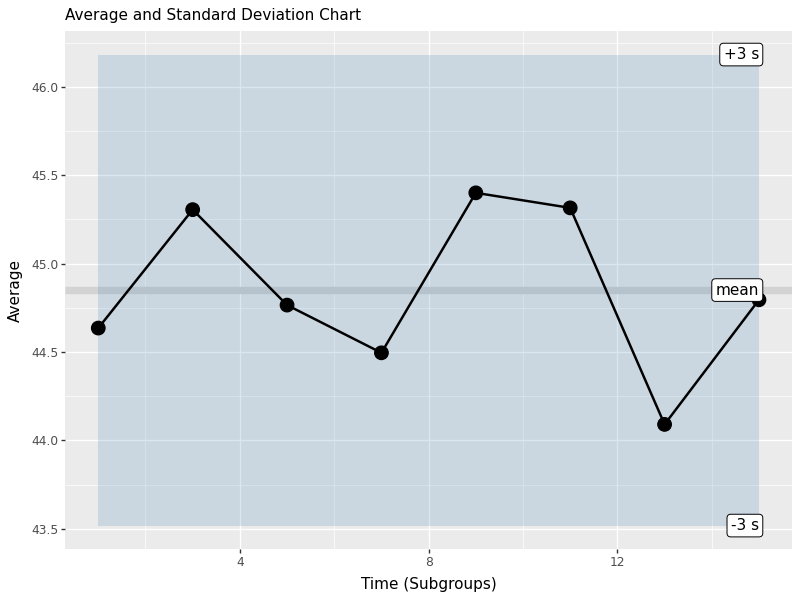
This control chart shows:
- mid line: grand mean (xbbar)
- control limits: upper and lower 3-sigma limits based on standard error
- personal points:mean of each subgroup plotted over time
- shaded area: control border area
Points outside the control limits or showing non-random patterns indicate that the process may be out of control and requires investigation.
learning check 1
Question
Prepare similar process overview chart for pH,
[View Answer!]
def ggprocess(x, y, xlab='Subgroup', ylab='Metric'):
import pandas as pd
from plotnine import ggplot, aes, geom_hline, geom_jitter, geom_boxplot, labs
d = pd.DataFrame({'x': x, 'y': y})
g = (ggplot(d, aes(x='x', y='y', group='x')) +
geom_hline(aes(yintercept=d['y'].mean()), color='lightgrey', size=3) +
geom_jitter(height=0, width=0.25) +
geom_boxplot() +
labs(x=xlab, y=ylab, subtitle='Process Overview'))
return g
ph_chart = ggprocess(water['time'], water['ph'])
# Save the plot
ph_chart.save('images/05_ph_chart.png', width=8, height=6, dpi=100)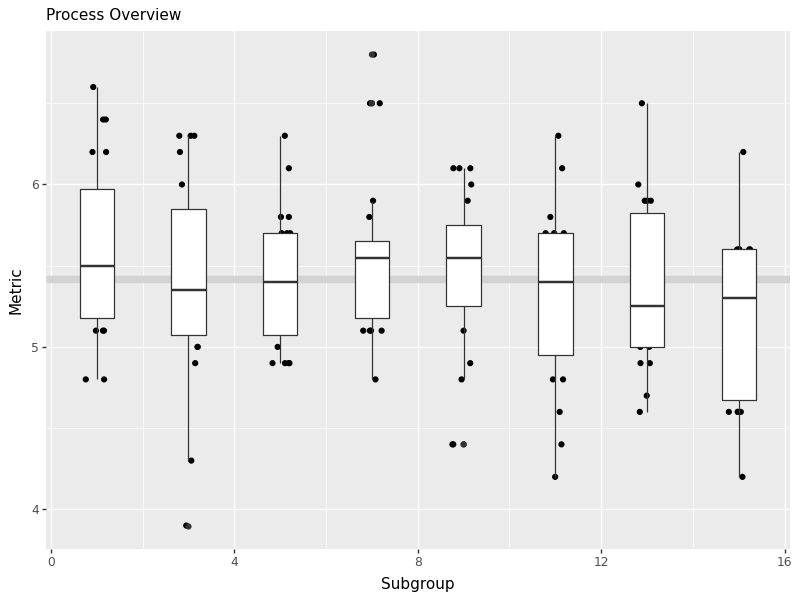
Moving Range Chart (n=1)
When we have individual measurements rather than subgroups, we use moving range charts. The moving range is the absolute difference between consecutive measurements, which helps us estimate process variation when we cannot calculate sub-group statistics.
indiv = water.iloc[[0,20,40,60,80,100,120,140]]
mr = (indiv['temp'].diff().abs().dropna())
mrbar = mr.mean()
import numpy as np
d2 = np.mean(np.abs(np.diff(np.random.normal(0,1,10000))))
sigma_s = mrbar / d2
se = sigma_s / (1**0.5)
upper = mrbar + 3*se
lower = 0istat = pd.DataFrame({'time': indiv['time'].iloc[1:], 'mr': mr, 'mrbar': mrbar, 'upper': upper, 'lower': lower})
mr_chart = (ggplot(istat, aes(x='time', y='mr')) +
geom_ribbon(aes(ymin='lower', ymax='upper'), fill='steelblue', alpha=0.25) +
geom_hline(aes(yintercept=mr.mean()), size=3, color='darkgrey') +
geom_line(size=1) + geom_point(size=5) +
labs(x='Time (Subgroup)', y='Moving Range', subtitle='Moving Range Chart'))
# Save the plot
mr_chart.save('images/05_moving_range_chart.png', width=8, height=6, dpi=100)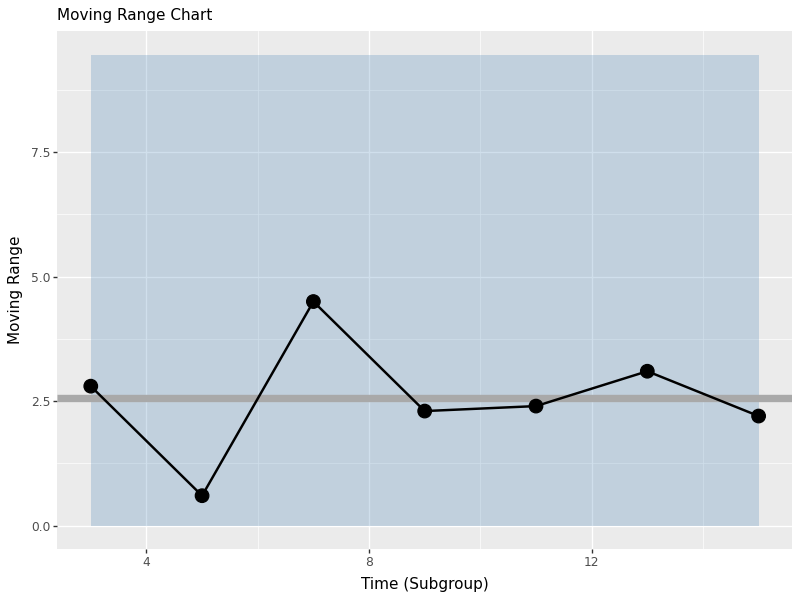
The moving range chart shows:
- mid line: Average Speed Limit (MRbar)
- upper control limit: Estimated process based on standard deviation
- lower control limit: Set to 0 (moving range cannot be negative)
- personal points:each moving range value
This chart helps us monitor process variation when we have individual measurements rather than subgroups.
conclusion
You have successfully created SPC visuals and statistics in Python: process overview, subgroup statistics, and moving range logic. These tools help us understand process behavior, identify when a process is in control or out of control, and make data-driven decisions about process improvements.
<a href